Current location: Home > NEWS > Industry news
NEWS
PRODUCTS
What are liquid biopsy techniques? A detailed explanation of circulating tumour cells and circulating tumour DNA
News source: Release time:[2022-07-20]
What is a liquid biopsy?
In the 10 Breakthrough Technologies 2015 list published by MIT Technology Review magazine, liquid biopsy technology was honoured on the list.
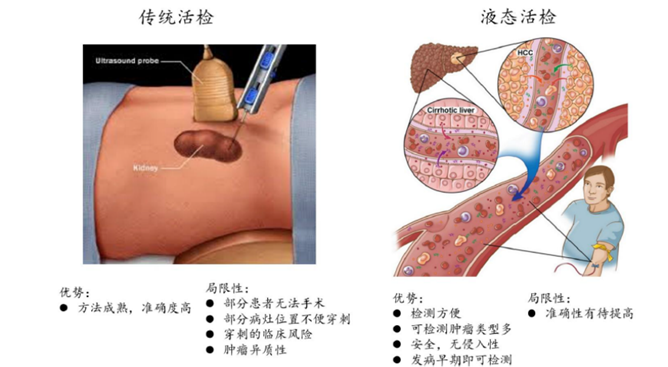
▲Comparison of traditional biopsy and liquid biopsy
Liquid biopsy refers to the analysis and diagnosis of cancer and other diseases by testing blood or other body fluids such as urine, saliva, pleural effusion, cerebrospinal fluid, etc. The patient's body fluid is mainly used as a tumor biopsy sample, and the objects captured and detected include circulating tumor cells (CTC), circulating tumor DNA (ctDNA) and exosomes in body fluids. Liquid biopsy is one of the hottest areas in the biomedical industry in recent years. Compared with standard tissue biopsy, liquid biopsy has some theoretical advantages: less trauma, reproducibility, real-time judgment of efficacy, dynamic adjustment of treatment decisions, etc.
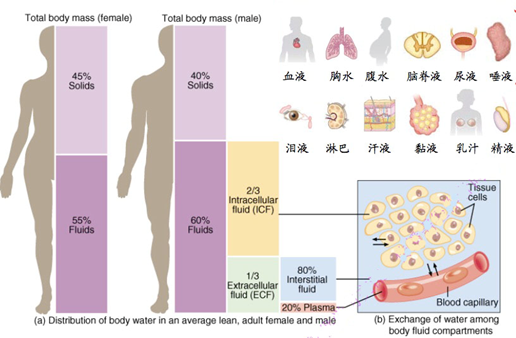
▲Sample type of liquid biopsy
The emergence of liquid biopsy technology marks another big step forward for mankind on the road to conquering cancer. Compared with traditional tissue biopsy, liquid biopsy has several advantages: First, liquid biopsy can be a substitute for areas that tissue biopsy cannot reach. Second, tissue biopsies can only reflect the information in the sample, while liquid biopsies can detect the overall condition of the patient. In addition, liquid biopsies primarily detect circulating tumour DNA (ctDNA), which typically travels from tumour tissue into blood vessels. As a result, liquid biopsies localize cancer cells more rapidly than relying on symptoms and images for diagnosis. Liquid biopsy technology is cost-effective. Through non-invasive sampling, it can not only greatly reduce the difficulty of cancer diagnosis and care, but also advance the time of cancer diagnosis and effectively prolong the survival period of patients.
#01
Circulating tumor cells (CTCs) and tumors
1)Circulating tumor cells (CTCs) are defined as those cancer cells that leave solid tumor lesions and enter the bloodstream. CTCs are not the only tumor derivatives in the circulation, but they contain a set of metastatic precursors that are essential to complete disease progression. Circulating tumor cells that enter the blood circulation contain complete genome, proteome and other biological information, and have the potential to develop into metastatic lesions, representing the aggressiveness of tumors.
However, technical conditions limit the isolation of live CTCs, mainly due to the difficulty in isolating CTCs in their rarity state compared to blood cells. But there is still no shortage of newly developed technologies that can both capture and isolate CTCs and test them at the molecular level for their suitability for clinical applications. Clinically, CTCs have demonstrated their value as predictive biomarkers of disease prognosis, and their suitability for other applications is being tested. Technically, it is generally necessary to enrich circulating tumour cells from massive blood cells and to distinguish malignant tumour cells from white blood cells and red blood cells.
2) In past explorations, a large number of specialized CTC isolation techniques have been developed that, in essence, can be divided into two broad categories: ① cells that capture CTCs based on specific antigen expression (positive selection); ② consume the All non-CTC cells (negative selection).
One of the classic cell-based (positive selection) techniques for CTCs is based on a two-step procedure: the first step involves centrifugation of the sample to eliminate plasma components, while CTCs are captured using anti-epithelial cell adhesion molecule (EpCAM) conjugated magnetic ferrofluid; In the second step, putative CTCs were further stained and identified using an anti-cytokeratin antibody, while contaminant white blood cells (WBCs) were identified by CD45 staining. Advances in single-cell sequencing technology are also facilitating the molecular analysis of cell samples of rare single CTCs.
3) Individual CTCs and CTC clusters. A CTC cluster is defined as a group of two or more CTCs with stable cell-cell junctions passing together through the blood. Compared with individual CTCs, CTC clusters displayed distinct gene expression profiles and dissemination patterns. Transcriptome analysis showed that CTC clusters retained epithelial features. Cell-cell junctions may play a key role in the formation and maintenance of the circulatory system. At present, related research on CTC clusters is also in full swing.
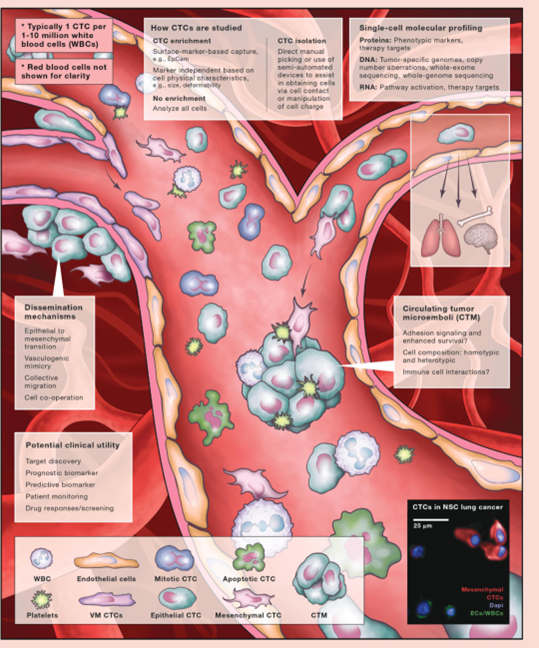
▲ CTCs and CTC clusters
#02
Circulating tumor DNA (ctDNA) and tumor
1)ctDNA is usually a DNA fragment actively secreted by tumor cells or released into the circulation during tumor cell apoptosis or necrosis, with a length of 132-145 bp and a short half-life (generally <2 h). ctDNA carries genetic features derived from tumor cells, such as gene mutation, methylation, amplification or rearrangement, and can be used as an important indicator for tumor screening, companion diagnosis, treatment efficacy evaluation, and prognostic risk stratification. Liquid biopsy is a minimally invasive method to visualize the sum of ctDNA at primary and secondary tumor sites. Not only can the amount of ctDNA be quantified, but genetic changes can also be identified. Specific mutations in genes have been identified in the plasma of several types of cancer patients, highlighting ctDNA as a possible cancer biomarker. However, achieving detectable ctDNA concentrations in body fluids is not trivial, and ctDNA fragments exhibit short half-lives.。
The expert consensus on clinical practice of ctDNA high-throughput sequencing in 2022 pointed out: With the rapid development of liquid biopsy, it is possible to use body fluids to analyze the molecular characteristics of patients, especially high-throughput based on circulating tumor DNA (ctDNA). Next generation sequencing (NGS) technology has been more and more widely used in clinic due to its advantages of non-invasive or minimally invasive, short detection time, ability to reflect intratumor and metastatic heterogeneity, and dynamic monitoring of therapeutic efficacy.
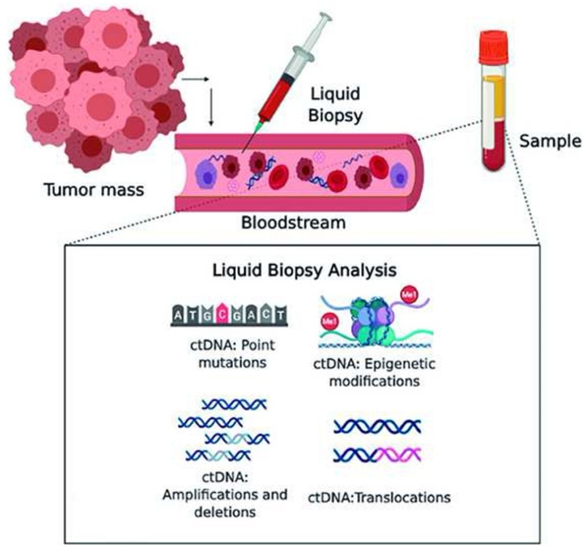
▲ctDNA levels are generally dynamic and influenced by multiple factors
ctDNA levels are generally dynamic and influenced by a variety of factors:
a.Factors such as tumor pathological tissue type, location, stage, and tumor burden can affect the release of ctDNA. ctDNA levels are generally lower in patients with tumors with lighter tumor burden, specific sites (eg, intracranial tumors) and specific histologies (eg, gliomas), and with lower levels of proliferation, apoptosis, and vascularization.
b.DNA interference from numerous other sources. Such as other normal cells or leukocyte-derived DNA, together with ctDNA is called cell free DNA (cfDNA).
c Gene mutation information carried by cfDNA produced by clonal hematopoietic cells may interfere with ctDNA assay results.
d. ctDNA has a short half-life and drug effects can also affect ctDNA content.
Compared with histological testing, ctDNA testing has the advantages of non-invasive or minimally invasive, repeated sampling, and short turnaround time for collection, processing and analysis reports. Plasma ctDNA also increased the detection rate of driver gene mutations compared with tissue biopsy alone. However, compared with tissue specimens, the content of ctDNA in peripheral blood is lower, which is prone to false negative results in clinical testing. Failure to use leukocytes as a control can also lead to false-positive results due to germline variation or clonal hematopoietic mutations. In addition, there are also differences in the amount of ctDNA released into the blood between different tumor patients or the same patient at different times, which also brings difficulties to the clinical detection and interpretation of ctDNA.
2 Application of ctDNA in companion diagnostics. Companion diagnostics (CDx) are considered to be an indispensable tool in tumor personalized medicine, which requires that the information provided by the test results is sufficient to ensure the safety and effectiveness of the corresponding therapeutic drugs. Patients who benefit from a specific treatment. The earliest clinical application of ctDNA detection as a CDx was epidermal growth factor receptor (EGFR) mutation detection, which was mainly used to identify patients with advanced NSCLC who may benefit from EGFR TKIs.

In the ENSURE study, ctDNA detected EGFR 19del and L858R mutations with a specificity of 98.2% and a sensitivity of 76.7%; and patients treated with erlotinib based on ctDNA results had significantly improved PFS compared with chemotherapy patients. Based on the results of this study, the FDA approved the first liquid biopsy CDx product, Cobas EGFR Mutation Test v2, in 2016, followed by the National Medical Products Administration (NMPA) in 2019 to approve the domestic launch of this CDx product. In the FLAURA study, the objective response rate (ORR) of ctDNA T790M-positive patients to osimertinib was consistent with that of patients with positive tumor biopsy. This has opened a wave of research hotspots in the detection of ctDNA in minimal residual disease (MRD) and homologous recombination deficiency (HRD).
3 Key points of ctDNA NGS detection technology. The middle stage of the analysis includes two steps: "wet experiment" and "dry experiment". "Wet experiment" includes nucleic acid extraction, primer or probe design and synthesis, library preparation, on-board sequencing, etc. The library preparation can choose hybridization capture or amplification. The quality control of this stage should make corresponding requirements for nucleic acid concentration, purity and fragment distribution in nucleic acid extraction and comply with the implementation. The "dry experiment" covers each step of the bioinformatics analysis process, and the quality control at this stage should be the minimum sequencing depth, average sequencing depth, coverage uniformity, guanine and cytosine (GC) content, base calling quality value, Comparing the quality value and on-target rate, etc., make corresponding requirements and comply with them. The post-analysis phase includes interpretation of the test report, discussion on the molecular tumor board (MTB), and genetic counseling.
Library Construction Strategies Targeted sequencing strategies for ctDNA detection using NGS methods are mainly divided into two types: hybrid capture-based NGS (hybrid capture NGS) and amplification-based NGS (amplification-based NGS). The hybrid capture-based targeted NGS method is a method in which the capture probe hybridizes to a specific region in the genome to obtain and analyze the sequence of the target region. This method has many operation steps, and the process is relatively complicated. It takes 1~2 days to complete the library construction and go on the computer. It can detect point mutations, small fragment insertions and deletions and copy number variations, as well as gene rearrangements at the DNA level, and some unknown fusions can be found. , the detection sensitivity can reach more than 95%. The probes used in the library construction of the target sequence capture method are designed with DNA probes and RNA probes, mainly for targeted enrichment of the target region, to ensure the complete enrichment of the sample target sequence, to improve the enrichment efficiency and the uniformity of coverage, and to effectively reduce the Sequencing cost. Amplicon-based targeted NGS approaches rely on multiple PCR amplification steps to enrich target sequences. During experiments, the actual hands-on time for amplification-based library preparation is usually shorter than for hybrid capture methods. Regardless of the method used, clear criteria for eligible libraries should be set for each assay.
Summarize
The emergence of liquid biopsy is the improvement of the existing tumor detection system. Liquid biopsy can detect traces of tumors in the body at an early stage, and is an important method for early screening of tumors. In addition, compared with tissue biopsy directly taken from tumor tissue, the material of liquid biopsy is derived from blood, which is less invasive and convenient for real-time monitoring. Positive effect is an important tumor diagnosis and treatment technology.总结
references
[1]. Expert consensus on clinical practice of ctDNA high-throughput sequencing (2022 edition)
[2].Next generation sequencing (NGS): a golden tool in forensic toolkit PMID: 27543959
[3].Clinical relevance of blood-based ctDNA analysis: mutation detection and beyond.PMCID: PMC7852556
[4].Tracking cancer progression: from circulating tumor cells to metastasis.PMCID: PMC7082968